Welcome to CircleBase
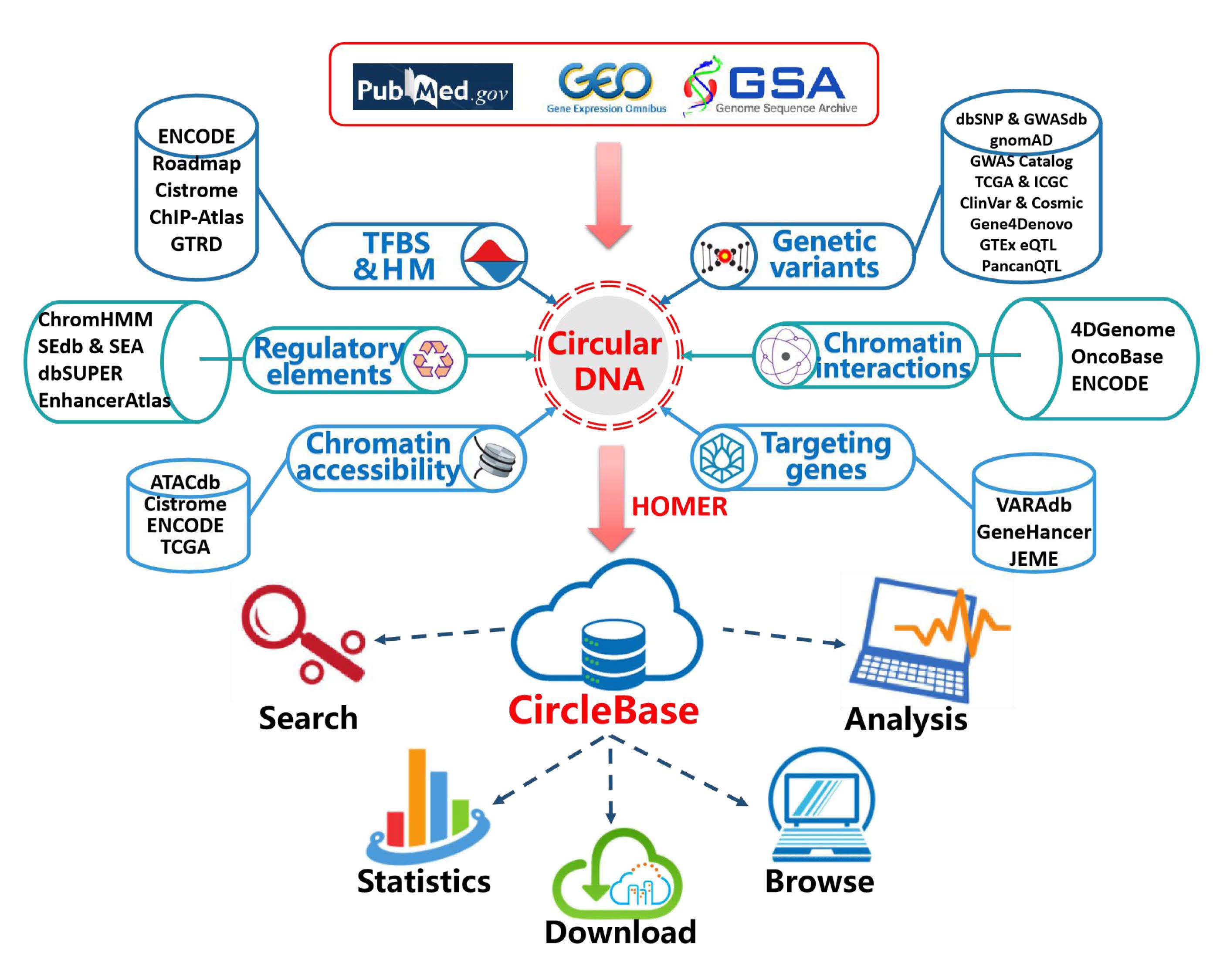
Here, we have developed a novel database, CircleBase, to provide interpretation of extrachromosomal circular DNAs (eccDNAs) which have been reported by computational analysis of sequencing data. CircleBase identifies putative functional eccDNAs by incorporating high-throughput experimental datasets from ENCODE and other sources, computational predictions and manual annotations. Briefly, CircleBase classifies the genomic and epigenomic data into six annotation sections, including 'targeting genes', 'epigenetic regulations', 'regulatory elements', 'chromatin accessibility', 'chromatin interactions', and 'genetic variants'. These data sources are combined into a powerful system to score eccDNAs and help interpreting the potential functions of eccDNA in the human genome. CircleBase provides a user-friendly interface for browsing, searching and downloading eccDNAs in various cell/tissue types. This platform will facilitate the identification of functional eccDNAs and help researchers to investigate the regulation networks between eccDNAs and genetic/epigenetic factors. We are dedicated to maintaining CircleBase and extending the range of organisms and cell types to keep the resource updated.
Workflow of CircleBase
Motivation of CircleBase
Extrachromosomal circular DNA (eccDNA) originates from chromosomal DNA but is independent of it. EccDNA is ubiquitous in various eukaryotes and plays multiple biological roles in different cell types. Researchers have known for decades that eccDNA occurs in tumor cells. However, comprehensive studies on their structure and function have returned relatively little useful information because of the limitations of available detection and analysis technology. Other researchers have exploited recently developed high-throughput sequencing technology and eccDNA identification methods, revealing strong associations between eccDNA and cancers. It indicates that eccDNAs are promising biomarkers for cancer diagnosis and prognosis and are, therefore, research hotspots. However, the number of human eccDNAs generated by the computational analysis of sequencing data has increased. Therefore, it has been challenging for researchers to compile available data to investigate their functions. In fact, the biological functions of most eccDNAs remain unknown and loss-of-function experiments are difficult to perform, as circular and linear chromosomes overlap in most cases. Moreover, the formation and regulation of most eccDNAs remain unclear and their targeting genes, genetic variants, and chromatin characteristics in various cell types have seldom been explored. Therefore, an integrated database and analysis platform is required for human eccDNAs. Herein, we build up an important analysis platform for exploring the biological function of eccDNA in the human genome and its application in cancer diagnosis and treatment.
Recent Updates
[3/8/2021] Finish scoring system for eccDNA prioritization and diffusion system to prioritize target genes.
[26/7/2021] The high-resolution chromatin interactions, clusters of transcription factor binding, somatic mutations, enhancers or super-enhancers and their predicted targets were illustrated in a circular ideogram layout by BioCircos.
[6/7/2021] Function enrichment analyses was performed using the R package clusterProfiler version 4.0.2.
[26/6/2021] Regulatory networks were constructed by combining enhancer-gene pairs from three sections including “Targeting genes”, “Regulatory elements” and “Chromatin interactions”.
[12/6/2021] Development of query system done.
[22/4/2021] Associate data from multiple sources and deposit it into the MySQL.
[2/11/2020] Data cleaning were done.
[27/9/2020] Retrieved data from multiple sources such as GEO, GSA, ENCODE, etc.
[5/1/2020] Website environment configuration, design and analysis method confirmation.